New brain atlas of transgenic mouse disease models shared via the Human Brain Project infrastructure
29 March 2019
Changing the expression of specific disease-related genes in animal models allows researchers to gain important insights into to molecular origins of neurodegenerative and other diseases of the brain in preclinical research.
In a recent Scientific Data paper, researchers from the University of Oslo and The Baylor College of Medicine in Houston present a novel online brain atlas of transgenic mouse lines that are used to generate mouse models of brain diseases, in which gene activity (so called “expression”) can be regulated by administration of the compound tetracycline. This allows researchers to investigate the effects of a transgene by reversibly turning its expression on or off in a controlled experimental setting. Changing the expression of specific disease-related genes in animal models allows researchers to gain important insights into to molecular origins of neurodegenerative and other diseases of the brain in preclinical research.
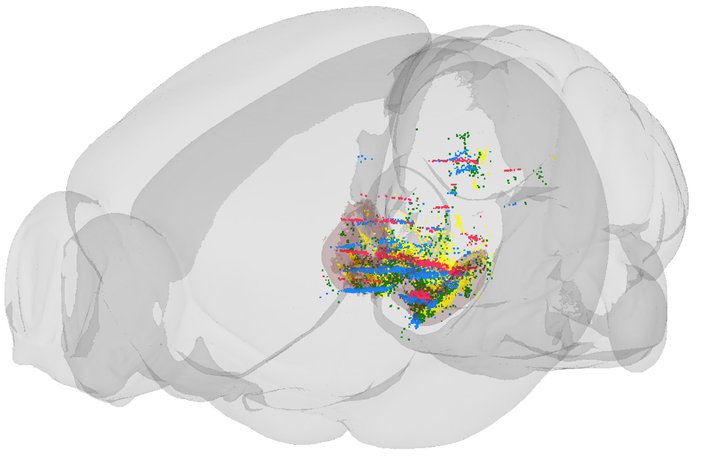 Three-dimensional visualization of where in the brain a regulatable gene promoter is expressed in four different transgenic mice. Reference to Lillehaug et al., 2019.
Three-dimensional visualization of where in the brain a regulatable gene promoter is expressed in four different transgenic mice. Reference to Lillehaug et al., 2019.
“This atlas shows us where in the brain each line controls expression of tetracycline-responsive transgenes, and therefore makes it easier to select the most appropriate conditional line for models of specific brain diseases”, explains Professor Joanna Jankowsky, who develops transgenic mouse models to unravel the molecular basis of Alzheimer’s disease. “By registering brain-wide series of histological section images to a common reference atlas, it becomes easier to compare where in the brain specific transgenes are expressed”, says PhD student Sveinung Lillehaug, who performed the anatomical analyses in the study.
The atlas is publicly shared via the research infrastructure established by the EU Human Brain Project. Professor Jan Bjaalie, HBP Infrastructure Operations Director, concludes that the project has been fruitful. – "This project is the first to share data with digital object identifiers (DOIs) issued from the Human Brain Project, Bjaalie says, “all data are assigned metadata, spatially defined in a reference atlas, and shared in accordance with the FAIR principles stating that data should be Findable, Accessible, Interoperable and Reuseable.
In the HBP, brain scientists and computing experts work together to handle the extremely large and heterogeneous datasets generated by modern neuroscience methods, and develop the technology and workflows for investigating the brain with High Performance Computing – both for the rodent and human brain. The resulting atlas and analytics tools are available to researchers around the world through their web browser. Dedicated support teams help researchers who want to use these ressources or integrate their data.
Project supervisor Professor Trygve Leergaard hopes that the community discovers the value of the collection, and encourages researchers who work with these conditional models to use the Human Brain Project pipeline to contribute their own data to the collection.
Original Publication:
Brain-wide distribution of reporter expression in five transgenic tetracycline-transactivator mouse lines. Sveinung Lillehaug, Michael J. Yetman, Maja A. Puchades, Martyna M. Checinska, Heidi Kleven, Joanna L. Jankowsky, Jan G. Bjaalie & Trygve B. Leergaard
Scientific Data volume 6, Article number: 190028 (2019)
https://www.nature.com/articles/sdata201928
See also:
Watch: Data integration and interactive use of HBP’s brain Atlas systems
The HBP atlas systems offer a way to integrate vast and heterogeneous neuroscience data into common frameworks and better understand the multilevel organization of the brain. A new video with HBP Infrastructure Operations Director Jan Bjaalie explains the key principles of HBP workflows used for registration of data into common reference atlases.
https://www.humanbrainproject.eu/en/follow-hbp/news/watch-data-integration-and-interactive-use-of-hbp-s-brain-atlas-systems/
Linking gene expression to human brain microstructure
24. 04. 2018 A team of researchers from the European Human Brain Project (HBP) has developed the JuBrain Gene Expression tool (JuGEx) that combines the benefits of a genetic and an anatomical atlas. Using JuGEx, the scientists found differentially expressed candidate genes for major depressive disorder in a disease-affected part of the brain.
https://www.humanbrainproject.eu/en/follow-hbp/news/linking-gene-expression-to-brain-microstructure/



