- Feature
Mapping at multiple scales – the most detailed atlas of the human brain
16 May 2023
The Human Brain Project has developed an atlas of the human brain with unprecedented detail and has made it freely available for everyone to browse online on the EBRAINS platform.
The human brain atlas can be compared to Google maps – for the brain instead of planet Earth. Just like Google maps, anyone can access the brain atlas using a simple web browser. Whereas an atlas of the world includes political, topographical or traffic information, the atlas of the human brain displays microstructure, connectivity and function. Zooming into the brain atlas, you can see cells instead of cities, and zooming out, you can see the borders of brain areas instead of countries.
When clicking on the name of a country in Google maps, or on an area in the brain atlas, a sidebar with additional information about the region pops up, and you can go beyond just looking at the maps and information. The brain atlas allows you to extract the underlying data to run an analysis or to put them into a simulation.
Like its counterpart, the brain atlas is dynamic, allowing continuous addition of new information. Via the Human Brain Project’s EBRAINS infrastructure, research groups from all over the world can integrate their own data into the living atlas, enabling the scientific community to work collaboratively on decoding the human brain.
Drawing borders
Brain mapping has a hundred-year tradition; yet, the concept of the HBP’s atlas is fundamentally new. Not just because of its digital and three-dimensional nature, but because it accounts for inter-subject variation. This is a crucial difference when comparing mapping the brain to mapping the world: while there is only one planet Earth, there are about eight billion human brains on it – and they are all slightly different. The question is: how different? And what do these differences mean for function?
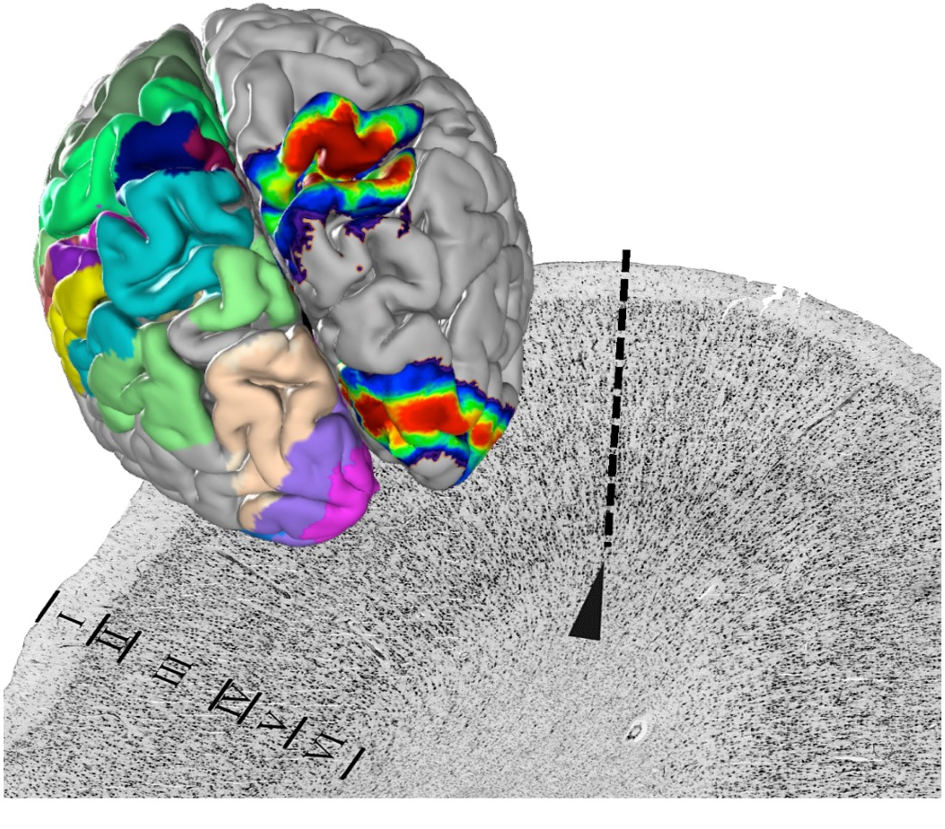
HBP researchers from Forschungszentrum Jülich and the Heinrich-Heine University Düsseldorf in Germany have stained the cell bodies of thousands of ultra-thin brain sections produced from 23 post mortem brains, digitised them and analysed their cytoarchitecture – the distribution, density and morphology of cells (Amunts et al. 2020). This microstructure of the brain reveals its parcellation into clearly distinguishable areas.

The team digitally reconstructed the mapped areas in a three-dimensional space and superimposed the maps of ten different brains for each area to generate probabilistic maps that show exactly how much the localization and size of an area varies from one individual to another.
The Julich Brain Atlas, which forms the centrepiece of the HBP’s Multilevel Human Brain Atlas, already contains more than 200 such probabilistic maps – including dozens of previously unmapped areas – and new ones are continuously added. Some of the most recent examples include maps of the insula, a large region that supports the integration of many functions including interoceptive, sensorimotor, cognitive and social-emotional processing (Quabs et al. 2022) and new areas of the anterior prefrontal cortex, a region that plays a major role in cognitive functions (Bruno et al. 2022).
Combining modalities
To understand a system as complex as the human brain, researchers need to combine insights from different levels of organisation. The HBP’s atlas enables just that – by supporting the integration of data from different scales and modalities into one common reference space. Modalities already integrated into the atlas include functional data generated by cognitive scientists from the French National Institute for Research in Digital Science and Technology (Inria) (Dadi et al. 2020) and density measurements of neurotransmitter receptors produced by researchers from Forschungszentrum Jülich (Palomero, Kedo et al., 2020).
Combining different modalities demonstrates that brain areas are more than just a system of structural parcellation: they have a physiological meaning as functional units of the brain that are distinct from one another on several levels. This is exemplified by a recent study showing how microstructural differences of brain areas correlate with distinct functions in visual-spatial orientation and associative memory (Stenger et al. 2022).
HBP researchers have also recently combined the analyses of the cytoarchitecture, the distribution of neurotransmitter receptors and the transcription of neurotransmitter receptor genes to uncover basic principles about the organisation of functional units of the brain (Zachlod et al. 2022). Each cytoarchitectonic area has a distinct pattern of receptor architecture and gene expression; however, a simple “mosaic” of brain areas does not explain how specific areas work together in an organised fashion, each playing a distinct role, resulting in cognitive functions. The researchers studied 15 cytoarchitectonic areas within the visual, auditory, somatosensory and motor systems and found that receptor distributions and gene expression change gradually along structural hierarchies within each functional system. In other words, they revealed that the changes occur in a systematic way in parallel with an increasing complexity of information processing.
Tracing connections
But how do distant parts of the brain work in concert to give rise to complex human perception and behaviour? To understand this, there is no getting around examining how individual cells and entire brain regions are connected with each other. To this end, the atlas includes connectivity data generated by HBP researchers from NeuroSpin in France in collaboration with a team at the University of Concepción in Chile (Guevara et al. 2017). The researchers have modelled fibre tracts at millimetre resolution based on diffusion MRI data from 78 healthy individuals. To study nerve fibres in post mortem brains at very high resolution, researchers at Forschungszentrum Jülich have developed a technique called three-dimensional polarised light imaging (3D-PLI).
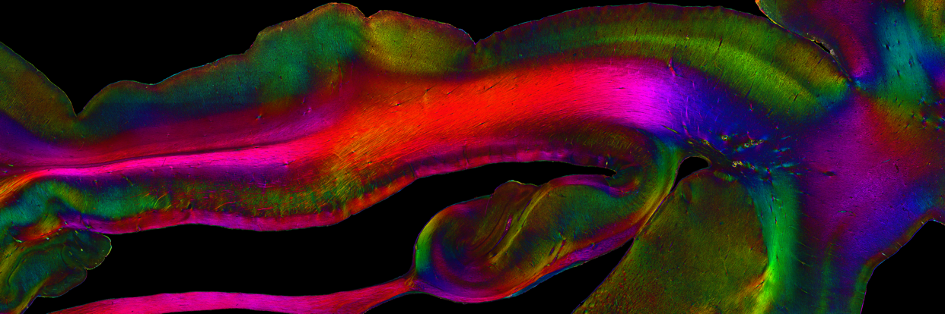
In order to bridge connectivity data from several spatial scales, HBP researchers from NeuroSpin, Jülich and the University of Florence have recently imaged one and the same tissue block from the human hippocampus using several different methods: anatomical and diffusion magnetic resonance imaging (aMRI and dMRI) 3D-PLI and two-photon fluorescence microscopy, respectively (Axer & Amunts 2022).
By integrating the resulting data from the different spatial scales into one common reference space within the atlas, the teams have provided crucial insights into the microstructural characteristics of complex fibre bundles.
Advancing brain medicine
The HBP’s atlas not only empowers researchers to study the healthy brain but has also become an important tool for better understanding brain disorders. Recently, researchers from Jülich, Düsseldorf and the Ernst von Bergmann Klinikum in Potsdam, Germany, have used the atlas to show in detail that in Parkinson’s disease the volumes of certain brain regions decrease over time in a specific pattern that is associated with clinical symptoms and largely coincides with the pattern described in Braak’s famous staging theory (Pieperhoff et al. 2022).
The atlas also contributes to directly improving medical treatments. For example, clinical researchers from Aix-Marseille University in France, are using the high-precision data of the atlas to optimise surgery of patients with epilepsy who don’t respond to pharmacological intervention. The researchers have developed personalised brain models to identify the areas where seizures emerge in a patient’s brain. A 400-patient clinical trial is currently ongoing with the aim of providing surgeons with a precise tool to help individual surgery decisions and improve outcomes (Wang et al. 2023). The HBP’s Multilevel Human Brain Atlas serves to enhance the accuracy of the method.
Text by Lisa Vincenz-Donnelly.
This text was first published in the booklet ‘Human Brain Project – A closer look at scientific advances’, which includes feature articles, interviews with leading researchers and spotlights on latest research and innovation. Read the full booklet here.
References
Amunts K, Mohlberg H, Bludau S, Zilles K (2020). Julich-Brain: A 3D probabilistic atlas of the human brain’s cytoarchitecture. Science 369(6506):988–992. doi: 10.1126/science.abb4588
Axer M, Amunts K (2022). Scale matters: The nested human connectome. Science 378(6619):500-504. doi: 10.1126/science.abq2599
Bruno A, Bludau S, Mohlberg H, Amunts K (2022). Cytoarchitecture, intersubject variability, and 3D mapping of four new areas of the human anterior prefrontal cortex. Front. Neuroanat. 16:915877. doi: 10.3389/fnana.2022.915877
Dadi K, Varoquaux G, Machlouzarides-Shalit A, Gorgolewski KJ, Wassermann D, Thirion B, Mensch A (2020). Fine-grain atlases of functional modes for fMRI analysis. NeuroImage 221:117126. doi: 10.1016/j.neuroimage.2020.117126
Guevara M, Román C, Houenou J, Duclap D, Poupon C, Mangin JF, Guevara P (2017). Reproducibility of superficial white matter tracts using diffusion-weighted imaging tractography. NeuroImage 147:703-725. doi: 10.1016/j.neuroimage.2016.11.066
Palomero-Gallagher N, Kedo O, Mohlberg H, Zilles K, Amunts K (2020). Multimodal mapping and analysis of the cyto- and receptorarchitecture of the human hippocampus. Brain Struct. Funct. 225(3):881-907. doi: 10.1007/s00429-019-02022-4
Pieperhoff P, Südmeyer M, Dinkelbach L, Hartmann CJ, Ferrea S, Moldovan AS, Minnerop M, Diaz Pier S, Schnitzler A, Amunts K (2022). Regional changes of brain structure during progression of idiopathic Parkinson's disease - A longitudinal study using deformation based morphometry. Cortex 151:188 210. doi: 10.1016/j.cortex.2022.03.009
Quabs J, Caspers S, Schöne C, Mohlberg H, Bludau S, Dickscheid T, Amunts K (2022). Cytoarchitecture, probability maps and segregation of the human insula. NeuroImage 260:119453. doi: 10.1016/j.neuroimage.2022.119453
Stenger S, Bludau S, Mohlberg H, Amunts K (2022). Cytoarchitectonic parcellation and functional characterization of four new areas in the caudal parahippocampal cortex. Brain Struct. Funct. 227(4):1439-1455. doi: 10.1007/s00429-021-02441-2
Wang HE, Woodman M, Triebkorn P, Lemarechal JD, Jha J, Dollomaja B, Vattikonda AN, Sip V, Medina Villalon S, Hashemi M, Guye M, Makhalova J, Bartolomei F, Jirsa V (2023). Delineating epileptogenic networks using brain imaging data and personalized modeling in drug-resistant epilepsy. Sci. Transl. Med. 15(680):eabp8982. doi: 10.1126/scitranslmed.abp8982
Zachlod D, Bludau S, Cichon S, Palomero-Gallagher N, Amunts K (2022). Combined analysis of cytoarchitectonic, molecular and transcriptomic patterns reveal differences in brain organization across human functional brain systems. NeuroImage 257:119286. doi: 10.1016/j.neuroimage.2022.119286



