- Paper Digest
How FAIR workflows help to combine models and methodologies in neuroscience
08 November 2022
Modelling in neuroscience is an essential complement to experiments in the efforts to understand the brain, and today a large ecosystem of different models exists. These describe different parts and processes in the brain, from subcellular details to large networks of interacting neurons, but are also developed using different methodologies, like a more data-driven or hypothesis-driven approach.
Hypothesis-driven modelling typically brings a question into focus so that a model is constructed to investigate a specific hypothesis. In data-driven modelling, on the other hand, model construction is informed by the computationally intensive use of data.
In a recent review paper, researchers of the Human Brain Project (HBP) argue that even though these different types of models can be useful, their value is greatly increased if they can be integrated to understand the same experimental data and phenomena at multiple scales or different levels of abstraction. They suggest that such integration can be achieved through the use of more standardized modelling workflows, and they use the FAIR concept (Findability, Accessibility, Interoperability and Reusability), originating from data management, to describe this.
The team of researchers from different institutions in Sweden, Switzerland, Finland, Lithuania and Germany, as well as India and the USA, point out that combining these models while validating them against experimental data is important to increase the understanding of the brain. However, integrating models that represent different biological scales and are built with different modelling philosophies requires a high degree of interoperability, transparency and reusability of both models and the workflows used to create them.
To solve this issue, the authors argue that applying the FAIR principles to models and software as well as the experimental data would help researchers to find, reuse, question, validate and extend published models, regardless of how they are implemented.
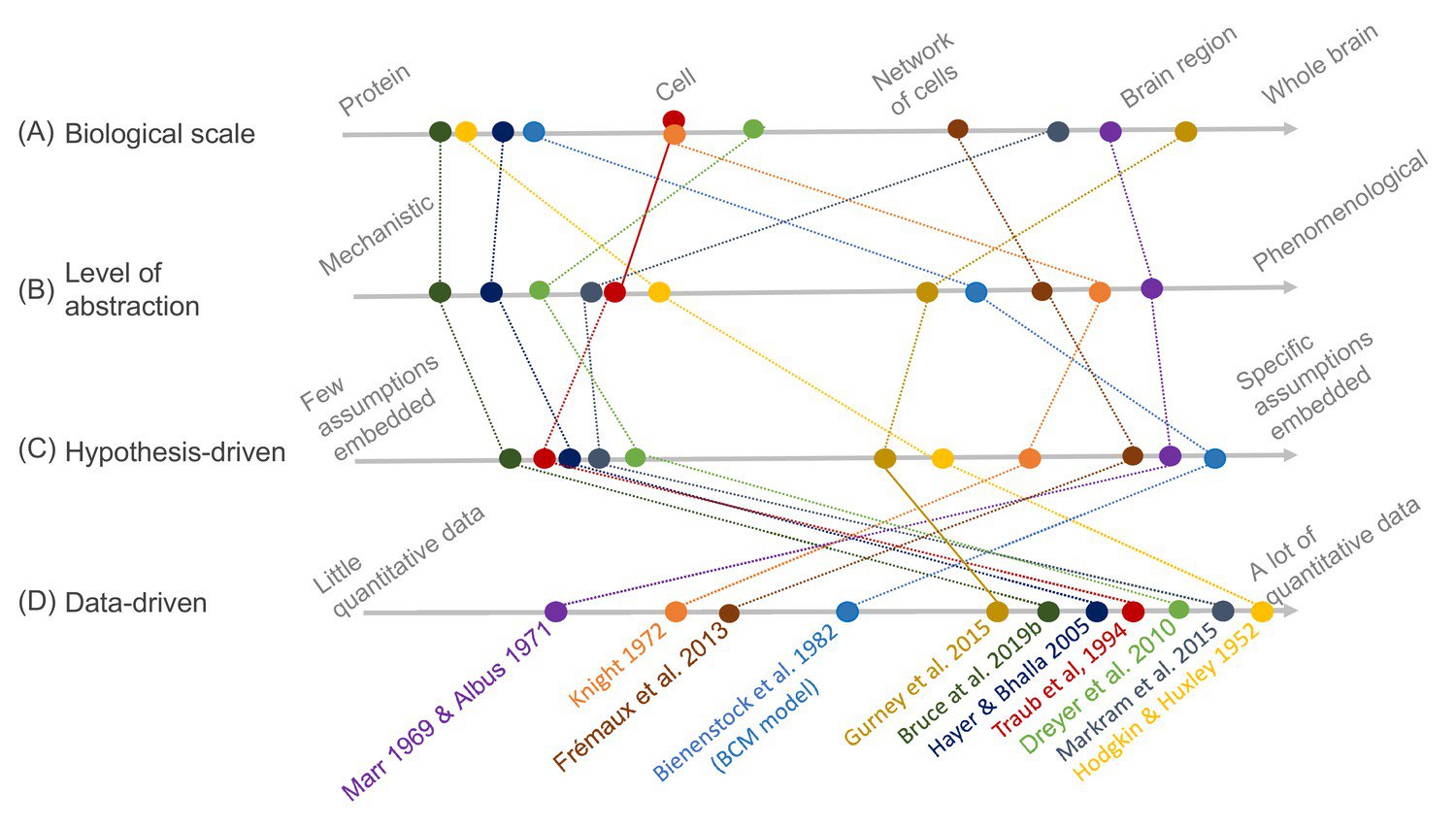
To illustrate this idea, the team used a synaptic plasticity model called the Bienenstock-Cooper-Munro (BCM) rule as a concrete example. The BCM rule was chosen by the researchers due to its long history (it was published in 1982, before the formulation of FAIR principles), different levels of abstraction and implementation at many scales.
The authors say that the history of models that implement the BCM rule demonstrates a clear progression from abstract, qualitative, and un-FAIR formulations to more mechanistic, quantitative, and FAIR formulations. This progression is a natural consequence of the accumulation of experimental data and nascent concepts of FAIR principles, they explain. In their review, the team considers how to improve neuroscience models further through FAIR workflows and data science-based approaches.
“We wanted to show how to combine data-driven and hypothesis-driven approaches. So, if you set up a model more focused on the research question, you can also use data science methods and test that question more efficiently”, explains Olivia Eriksson, researcher at KTH Royal Institute of Technology and first author of the paper.
The other goal was to assemble a knowledge resource for FAIR modelling. In the paper the authors have listed numerous examples of different databases, file formats, simulators and other modelling software, which can be combined to workflows, that are starting to align to the FAIR principles. They note however that the exact form of such workflows is an ongoing discussion within the neuroscience community.
The study benefited from the combined expertise of researchers with knowledge in different fields. “I think this paper would have been impossible to write if it was only one group. The brain is very complex and all these researchers are experts on different model types, software or methodologies; this combination of people made this work possible”, says Eriksson.
Text by Helen Mendes
Reference:
Olivia Eriksson, Upinder Singh Bhalla, Kim T Blackwell, Sharon M Crook, Daniel Keller, Andrei Kramer, Marja-Leena Linne, Ausra Saudargienė, Rebecca C Wade, Jeanette Hellgren Kotaleski (2022) Combining hypothesis- and data-driven neuroscience modeling in FAIR workflows eLife 11:e69013. https://doi.org/10.7554/eLife.69013



