- Press Release
HBP scientists have simulated how the Parkinson’s brain responds to deep stimulation at multiple scales
22 June 2022
Researchers of the Human Brain Project have created the first multiscale model of how a Parkinson’s brain responds to deep brain stimulation. Deep brain stimulation is a common method to treat Parkinson's symptoms, but currently outcomes of the invasive treatment are hard to predict. Modeling and simulating electric stimulation across multiple levels of brain networks can help clinicians “preview” their effects and plan therapies accordingly.
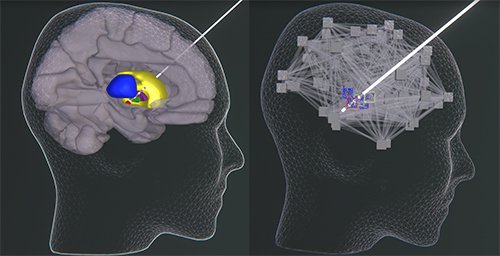
First clinical use case of multiscale co-simulation of the human brain applied for in silico optimization of deep brain stimulation in Parkinson’s disease. The Basal-ganglia-thalamus circuit is simulated at microscopic scale of spiking neurons while the remaining brain is being simulated at coraser scale of mean field models. Upon virtual deep brain stimulation spike trains of subcortical nodes and mean field activity in the remaining brain is being monitored.
The study marks the first published case of a multiscale co-simulation of the human brain applied for a clinical use case, and the methodology is already openly available on the EBRAINS platform, the digital research infrastructure developed by the Human Brain Project.
“Among the aims of HBP is the implementation of brain models at multiple scales, capable of showing high detail neuronal activity inserted into a larger context” explains Petra Ritter of Berlin Institute of Health at Charité, senior author of the study. “We previously described the methodology for doing this, and are now able to accomplish it in EBRAINS and identified a first use case: deep brain stimulation in Parkinson’s disease”.
Monitoring what happens during Parkinson’s at multiple scales is very useful in understanding the brain dynamics of the patient and implementing therapeutic deep brain stimulation. However, multiscale models are very rare and normally neuroscientists focus either to a small area in high detail, or on larger models that look at the mean field activity of the whole brain. “In the case of Parkinson's, you have small subcortical nuclei where you can look at the activity of single neurons not only in space, but also in time, monitoring the precise timing of spikes. You can also model plasticity effects that happen during stimulation of those areas” says Ritter. “But you would miss valuable information about what happens at the whole brain scale, which we can instead capture in The Virtual Brain, an open-source platform for constructing and simulating personalised whole-brain network models”.
The team used microcircuit models of basal ganglia and thalamus areas. This network was already created and optimized to work at small scales by a team lead by Fred Hamker from the Department of Computer Science, Chemnitz University of Technology in Germany, who is a co-author of the study. “We plugged the whole network into TVB, first making sure that it had the same behaviour of the original one” says Ritter. “We then constructed two multi scale models, one for a Parkinson’s patient, the other for a healthy control, based on measured brain data for both models”. The researchers then compared first differences in resting state behavior of the healthy and the Parkinson’s brain model. Next, they started programming and simulating different targeted deep brain stimulations that are frequently done in vivo, in a clinical setting.
Interestingly, the researchers found that the firing rate of the subcortical regions and specifically of parts of the thalamus have a decreased rate in the Parkinson’s patient’s model during resting state – an observation that has also been reported for real Parkinson’s patients’ brains. This drop of firing rate could be normalised through in-silico deep brain stimulation. “However, this also caused an effect at the cortical scale, resulting in differential activity of the motor cortex depending on the stimulus. Some effects were potentially less beneficial for the patient” says Ritter. “This whole brain effect would not have been noticed by only looking at the microscale, but became apparent when looking at multiple scales at the same time”.
With the new approach, it is possible to both monitor some structures in high detail (not only spatially, but also at a temporal scale) and experience the global effect of the simulation, concludes Ritter. “This has the potential to be translated into the clinical setting, improving prediction and personalisation when performing deep brain stimulation. The next step is to introduce plasticity rules into the system and see which other phenomena we are capable of capturing at the multiscale”.
Text by Roberto Inchingolo
Read the full paper: Jil M.Meier, Dionysios Perdikis, André Blickensdörfer, Leon Stefanovsky, Qin Liu, Oliver Maith, Helge Ü.Dinkelbach, Javier Baladron, Fred H.Hamker, Petra Ritter, Virtual deep brain stimulation: Multiscale co-simulation of a spiking basal ganglia model and a whole-brain mean-field model with The Virtual Brain, Experimental Neurology, Volume 354, August 2022, https://doi.org/10.1016/j.expneurol.2022.114111
Media contact
Peter Zekert
press@humanbrainproject.eu



