Cerebellum
Resources
The cerebellar neuron models can be found on the Brain Simulation Platform.
In detail:
- a dedicated use case called "Custom Axon Cerebellar Granule cell simulation" containing the code to build up the axon with BluePyOpt
- the code of the new granule cell models (Masoli et al.2019)
- the code for automatic fitting of the mouse mono compartmental granule cell model (Masoli et al., 2017)
- the code for automatic fitting of the mouse multi compartmental granule cell model (updated from Dover et al., 2016).
The following models can also be launched from the Collaboratory producing optimized granule cells models.
- the code of the mouse Purkinje cell model (derived from Masoli et al., 2015; Masoli and D'Angelo, 2017)
- the code of the mouse Stellate cell model (Rizza, Locatelli, Masoli, Munoz, D'Angelo, in preparation).
- the code of the mouse Golgi cell model (Masoli, Locatelli, Munoz, D'Angelo, in preparation).
- the first scaffold model of the mouse cerebellum, cell placement and connectome (Casellato, Casali, De Schepper, Geminiani, Antonietti, Medini, D'Angelo).
- simulation tests using scaffold model hosting cell-specific point neurons or morphologically multi-compartmental neurons (Casellato, Casali, De Schepper, Geminiani, Antonietti, Medini, D'Angelo).
- a dedicated pipeline for data communication with the NIP and for the use of HPC resources in running these models (PRACE cerebellum).
You can access them via the Cerebellum Collab.
If you do not have an account to access EBRAINS, please email
Publications
A list of recent and relevant publications:
-
Stefano Masoli , Marialuisa Tognolina , Umberto Laforenza , Francesco Moccia , Egidio D’Angelo . Parameter tuning differentiates granule cell subtypes enriching transmission properties at the cerebellum input stage. Nature Communications Biology 3, 222 (2020). DOI: 10.1038/s42003-020-0953-x.
-
Casali S, Marenzi E, Medini C, Casellato C, D’Angelo E, “Reconstruction and Simulation of a Scaffold Model of the Cerebellar Network” Frontiers in Neuroinformatics Vol 13, pags 37; 19/05/2019; doi: 10.3389/fninf.2019.00037
-
Geminiani A, Casellato C, D’Angelo E, Pedrocchi A. “Dynamics in Olivocerebellar Neurons Represented With Extended-Generalized Leaky Integrate and Fire Models” Front Comput Neurosci. 2019 Jun 6;13:35. doi: 10.3389/fncom.2019.00035
-
Geminiani A, Pedrocchi A, D’Angelo E, Casellato C, Response Dynamics in an Olivocerebellar Spiking Neural Network With Non-linear Neuron Properties. Front Comput Neurosci. 2019 Oct 1;13:68. doi: 10.3389/fncom.2019.00068
-
Casali S, Tognolina M, D’Angelo E, Cellular-resolution mapping uncovers spatial adaptive filtering at the cerebellum input stage, bioRxiv 2020.03.14.991794; doi: 10.1101/2020.03.14.991794
-
Masoli S, Tognolina M, Laforenza U, Moccia F, D’Angelo E, Parameter tuning differentiates granule cell subtypes enriching the repertoire of retransmission properties at the cerebellum input stage, bioRxiv 638247; doi: 10.1101/638247
__________________________
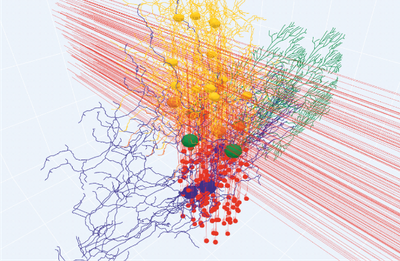 The cerebellum is a major neural structure that contains about half of the brain neurons with specific microcircuit organization and plasticity dynamics. The cerebellum plays an essential role in numerous cerebrocortical loops involved in sensori-motor control, cognition and emotion. A detailed model of the cerebellum is needed, not just to understand inherent microcircuit processing but also to implement large-scale simulations in virtual brains, robotic control systems and neuromorphic supercomputing architectures. Fundamental information on the role and mechanisms of cerebellum can be derived from multiscale simulations reconnecting microcircuit spatio-temporal dynamics to molecular and cellular properties of neurons and synapses.
The cerebellum is a major neural structure that contains about half of the brain neurons with specific microcircuit organization and plasticity dynamics. The cerebellum plays an essential role in numerous cerebrocortical loops involved in sensori-motor control, cognition and emotion. A detailed model of the cerebellum is needed, not just to understand inherent microcircuit processing but also to implement large-scale simulations in virtual brains, robotic control systems and neuromorphic supercomputing architectures. Fundamental information on the role and mechanisms of cerebellum can be derived from multiscale simulations reconnecting microcircuit spatio-temporal dynamics to molecular and cellular properties of neurons and synapses.What makes the cerebellum special?
What are the specific questions we want to address in the HBP?
The cerebellar microcircuit has been the workbench for theoretical and computational modelling since the beginning of neuroscientific research. The regular neural architecture of the cerebellum inspired different solutions to the long-standing issue of how its circuitry could control motor learning and coordination. Computational modelling in HBP therefore plays a major role in developing new theories about cerebellum and brain functioning.
What is our specific take?
We have developed a scaffold model of the cerebellar network along with detailed models of its neurons and synapses. These models are critical for implementing the Brain Simulation Platform and for explaining experimental data. This multiscale modelling strategy aims at the generation of the first full realistic model of the cerebellar network. The cerebellar model, in collaboration with the partnering projects "cerebNEST", "VM-cerebNEST", and "SPINNcer", is also going to be simplified and used in the Neurorobotics Platform and in the Neuromorphic computing platform to generate novel controllers and computational systems.
Roadmap
In March 2020, as part of the Brain Simulation Platform (BSP), we will complete the release of the fundamental set of detailed multicompartment cerebellar neuron models, along with their simplified single-point version, and the full cerebellar scaffold including cell placement and connectivity.
The aim was to build on the scaffold model developed in the previous phase in order to complete the missing microcircuit components, to update the mechanisms at the basis of these models according to the evolving physiological and anatomical discoveries, to embed subcellular components (such as synaptic plasticity mechanisms) into the scaffold model, to simplify the latter and embed it into large-scale brain loops.
These models can be used to investigate multiscale cerebellar circuit functions and for applications to neurorobotics, neuromorphic hardware and The Virtual Brain. Moreover, these models serve to aggregate the cerebellum modeling community around HBP and the future EBRAINS.
The models are developed using pyNEURON, pyNEST, TVB, (ARBOR in progress), SpiNNaker simulators and are accessible from the Brain Simulation Platform and Neurorobotic Platform.
Who's involved?
Building a model of cerebellum from experimental data will improve our understanding of the cerebellar and brain function. It requires a critical mass of people with differentiated expertise, from neurophysiology to biomedicine, neuroengineering and biophysics.
The following people are driving this effort:
Egidio D’Angelo, University of Pavia
Claudia Casellato, University of Pavia
Claudia Gandini Wheeler Kingshot, University of Pavia and UCL Institute of Neurology, London
Fulvia Palesi, University of Pavia
Simona Tritto, University of Pavia
Stefano Masoli, University of Pavia
Martina Rizza, University of Pavia
Stefano Casali, University of Pavia
Alberto Antonietti, University of Pavia
Alice Geminiani, University of Pavia
Alessandra Ottaviani, University of Pavia
Robin De Schepper, University of Pavia
Edoardo Negri, University of Pavia
Roberta Lorenzi, University of Pavia
Lisa Mapelli, Francesca Prestori, Marialuisa Tognolina, Teresa Soda, Teresa Sorbo, Giuseppe Gagliano, Ileana Montagna, Anita Monteverdi, and Danina Di Domenico are providing the critical data required for the model reconstruction and validation.
Alessandra Pedrocchi and Alessandra Trapani are contributing form cerebNEST.
What's in it for me?
Model use: in the Cerebellum Collab you will find the Data Centre with detailed information about the cerebellar network, the neurons and the connectivity rules.
Results of single cell models derived from optimization procedures can be found in:
- the Single Cell Optimisation (Purkinje cells, and granule cells): currently, only the granule cell optimisation procedure code is available
- the Granular Cell synthesis: a Jupyter notebook with python code for connecting granule cell dendrites to mossy fibre terminals is available
- the Circuit Building Pipeline: python code for the reconstruction of a network volume is available in a Jupyter notebook
- live papers: (1) Geminiani et al., 2019; (2) Casali et al., 2019; (3) Masoli et al., 2019
- Cerebellum showcase on the NRP
- Tutorials and videos of the Hackathon on cerebellum modelling
You can simply go to the Brain Simulation Platform and launch one of the use cases. Alternatively, you can download data and models as indicated in the above Resource box.
Participate in community modelling: we would be happy to hear from you if you would like to get involved and contribute to our community effort. For all information, please contact Simona Tritto.



