Basal Ganglia
Resources
- The first version of the mouse striatal microcircuit, the input stage of the basal ganglia, is implemented (LivePaper Hjorth et al., 2020; see also Jupyter notebook).
- Online use case on single cell building (BSP, Optimise a striatal fast-spiking interneuron).
- Online use cases on validation of the basal ganglia models (BSP, Basal Ganglia Single Cell Validation, Basal Ganglia Morphology Validation, Basal Ganglia Population Morphology Validation).
- Single cell model of striatal dSPN (MSN D1) (ModelDB, Lindroos et al., 2018).
- Experimental electrophysiological data for single neurons: SPN, FS, LTS and ChIN (Knowledge Graph at EBRAINS.eu).
- Morphological reconstructions of single neurons: SPN, FS, LTS and ChIN (Knowledge Graph at EBRAINS.eu).
- Electro-physiological feature extraction example (Collab, Jupyter notebook).
- Single cell optimisations of individual neurons: dSPN, (MSN D1), iSPN (MSN D2), FS, LTS, ChIN (LivePaper Hjorth et al., 2020).
Publications
A list of recent and relevant publications:
- Hjorth, J., Kozlov, A., Carannante, I., Frost Nylén, J., Lindroos, R., . . . Grillner, S. (2020). The microcircuits of striatum in silico. Proc Natl Acad Sci USA.
- Lindroos, R., Dorst, M. C., Du, K., Filipovic, M., Keller, D., Ketzef, M., . . . Hellgren Kotaleski, J. (2018). Basal Ganglia Neuromodulation Over Multiple Temporal and Structural Scales-Simulations of Direct Pathway MSNs Investigate the Fast Onset of Dopaminergic Effects and Predict the Role of Kv4.2. Front Neural Circuits.
- Du, K., Wu, Y. W., Lindroos, R., Liu, Y., Rozsa, B., Katona, G., . . . Kotaleski, J. H. (2017). Cell-type-specific inhibition of the dendritic plateau potential in striatal spiny projection neurons. Proc Natl Acad Sci USA.
- Grillner, S., von Twickel, A., & Robertson, B. (2017). The blueprint of the vertebrate forebrain - With special reference to the habenulae. Sem Cell Dev Biol.
- Grillner, S., & Robertson, B. (2015). The basal ganglia downstream control of brainstem motor centres--an evolutionarily conserved strategy. Curr Opin Neurobiol.
____________________________
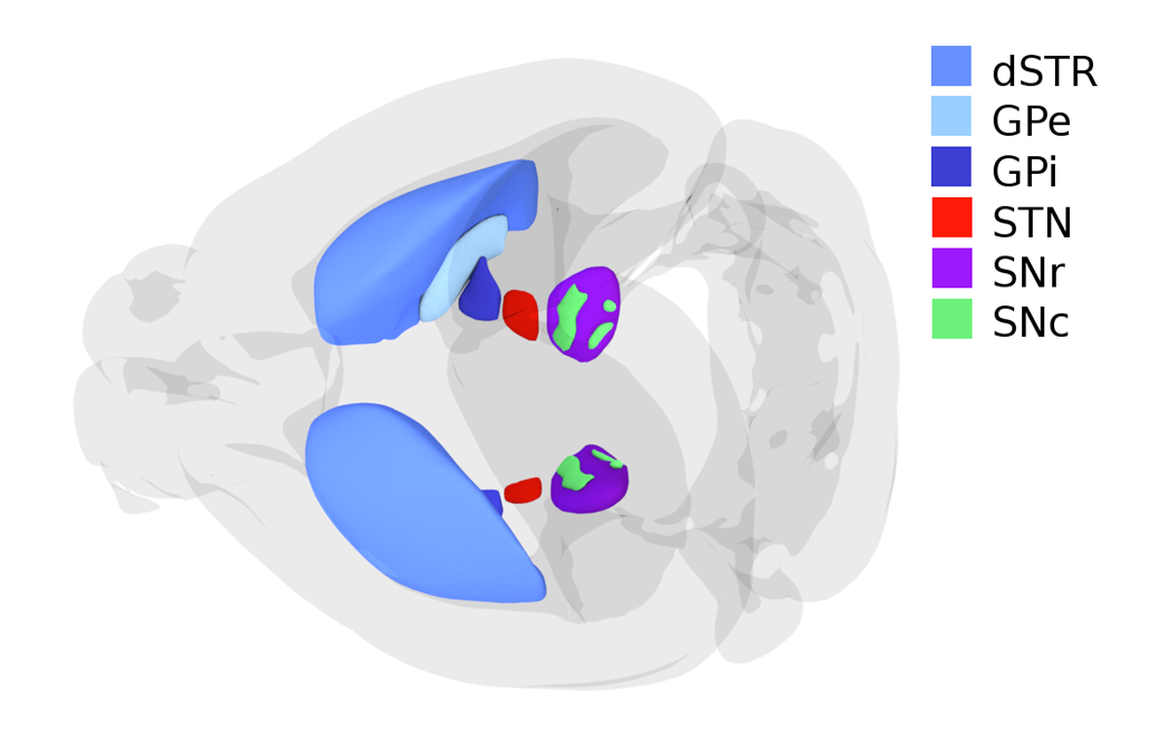
The basal ganglia is an evolutionarily conserved key structure for the control of action (Grillner and Robertson, 2015) we want to address in HBP. It plays a crucial role in decision-making and motor control. The output of the basal ganglia consists of tonically active GABAergic neurons, a proportion of which project to different brainstem centres and another part projects to the thalamus and back to the cortex. The target areas are tonically inhibited under resting conditions. These centres will be disinhibited when called into action.
What makes the basal ganglia special?
The basal ganglia is a highly interconnected structure within the brain which includes multiple distinct nuclei. The detailed structure of these nuclei at the meso- and micro-circuit levels is largely unknown and is a subject of intensive research. It exhibits a great degree of plasticity being a target of neuromodulatory systems (dopamine, serotonin and histamine). It also plays an important role in the regulation of the midbrain dopamine system, which is involved in reward, disappointment, aversion, etc.
What are the specific questions we want to address in the HBP?
The model will provide a ‘scaffold’ for a first data-driven model of the whole basal ganglia, including selected downstream motor centres and feedback via the thalamus to the cortex. The model will also provide a basis for investigations of the role of the basal ganglia in action selection and motor learning. The important question is to understand how a decision is made when provided with multiple conflicting alternative actions that could be taken? How is the correct decision learnt and facilitated in future tasks? What is the role of neuromodulators in shaping the dynamics of the basal ganglia network?
What is our specific take?
A multi-pronged strategy, culminating in a scaffold detailed computer model of a rodent basal ganglia, starting with the striatum.
The simulation platform developed initially for the striatal microcircuit, will be expanded gradually to incorporate other nuclei. Spatial inhomogeneity based on the Allen Brain gene expression data will be taken into account. Special attention will be paid to the implementation of the cortical and thalamic afferent projections, as well as to modelling of plasticity- and modulation-related dynamic changes in the circuitry.
Who's involved?
Building a model of the basal ganglia from sparse, fragmented data and improving our understanding of the basal ganglia function requires a critical mass of experimental data and expertise.
The following people and their teams are driving this effort:
Sten Grillner (Professor), Department of Neurophysiology, Karolinska Institute, Sweden
Jeanette Hellgren Kotaleski (Professor), Science for Life Laboratory, KTH Royal Institute of Technology, Sweden
Alexander Kozlov (PhD), Science for Life Laboratory, KTH Royal Institute of Technology, Sweden
Johannes Hjorth (PhD), Science for Life Laboratory, KTH Royal Institute of Technology, Sweden
Ilaria Carannante (PhD student), Science for Life Laboratory, KTH Royal Institute of Technology, Sweden
Johanna Frost-Nylén (PhD student), Department of Neurophysiology, Karolinska Institute, Sweden
Benefits to the community
Model use: If you would like to use some of the results of this work (e.g. take a striatal projection neuron for a spin, create your own variant thereof, analyse a striatal network, etc.) you can simply go to the Brain Simulation Platform and launch one of the respective use cases. Alternatively, you can download the data and models as indicated under Resources.



